The CUPE 3903 labour disruption has now ended. Courses and related academic activities that were suspended as a result of the strike will resume during the Remediation Period, which begins on Monday, April 22, 2024. Instructors will contact students with details about their courses. For more information please visit the labour disruption website.
Fostering discovery. Engaging community.
Inspiring futures.
The Faculty of Science is a hub of research and teaching excellence, fostering scientific discovery and tackling global challenges to create positive change in our world.
Explore York Science
Upcoming Events
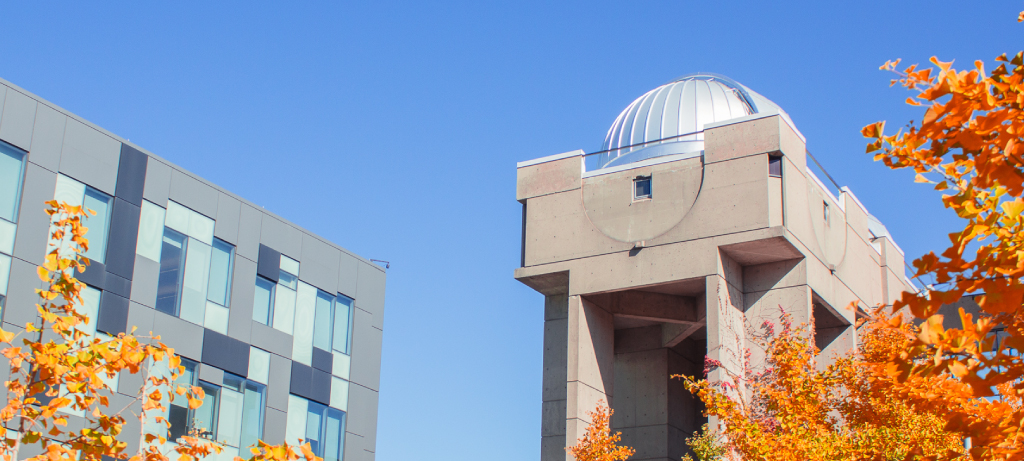

Allan I. Carswell Astronomical Observatory
Located on the York University Keele campus, the Allan I. Carswell Observatory supports student learning and research in astronomy and is a hub for public engagement and outreach. The Observatory offers a variety of free programming for the public.
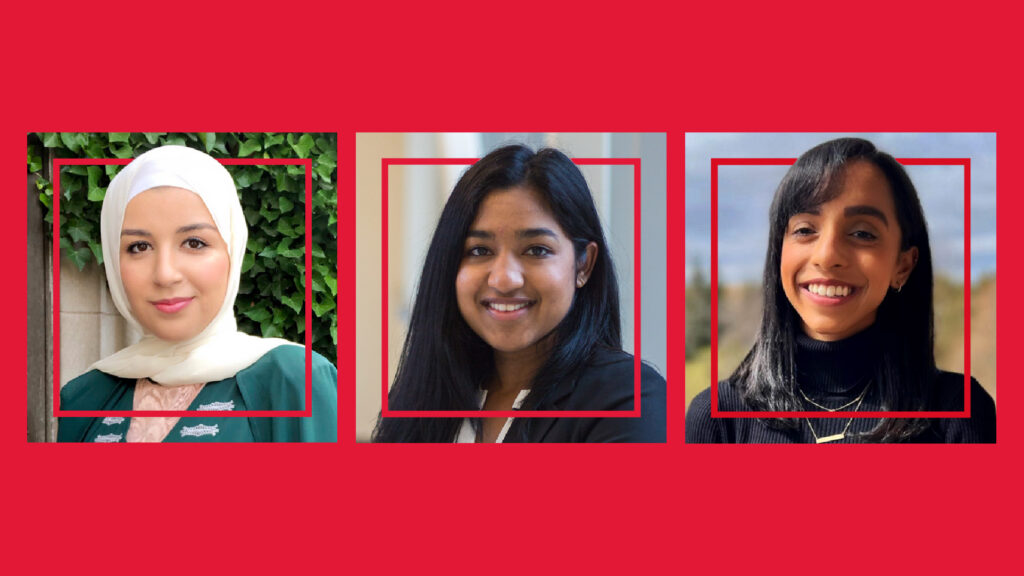

Three Faculty of Science alumni – Batool Barodi, Clarelle Gonsalves, and Shalini Iyer – were among the outstanding graduates announced as part of York University’s Top 30 Alumni Under 30 for 2023.

United Nations Sustainable Development Goals
The Faculty of Science rises to the York University-wide challenge to contribute to the United Nations Sustainable Development Goals (UN SDGs). Learn more about what we are doing to tackle these goals.